In this Article
- Section 01: PCR Basics: Understanding the Core Reaction
- Section 2: Building the Perfect Cocktail: A Deep Dive into PCR Reagents and Setup
- Section 3: The Engine of Amplification: Selecting the Right PCR Enzymes
- Section 4: Fine-Tuning the Machine: Optimizing PCR Cycling Parameters
- Section 5: Diverse Applications and Methods: Putting PCR to Work
- Section 6: Workflow Efficiency and Quality: Choosing the Best PCR Kits
- Section 7: The Practical Lab Fix: Your PCR Troubleshooting Guide
All of the products listed in AAA Biotech’s catalog are strictly for research-use only (RUO).

Summary/Learning
- PCR’s accuracy depends on precise reagent and enzyme selection, especially magnesium concentration and enzyme fidelity.
- Heat-stable polymerases automate workflow, replacing laborious manual enzyme addition.
- Primer design and concentration drive specificity, while template quality influences yield.
- MgCl₂ is vital: higher levels increase yield but reduce specificity.
- High-fidelity enzymes offer lower error rates but may reduce processivity.
- Hot Start technology prevents non-specific amplification during setup.
- Master mixes improve reproducibility and save time.
- Troubleshooting targets: primer annealing temperature, MgCl, template, and cycle count.
The Polymerase Chain Reaction (PCR) is often called the molecular photocopy machine because of its capacity to exponentially amplify specific regions of DNA.
While the underlying concept is ingeniously simple - copy, copy, copy - achieving consistent, high-yield results requires a deep and nuanced understanding of the core chemistry. This begins with the Principles of PCR and the precise roles played by every ingredient in the reaction mixture.
Understanding the individual PCR reagents, enzymes, and how to troubleshoot, leads to their being a core piece of the foundation of modern molecular biology.
Section 01: PCR Basics: Understanding the Core Reaction
1.1. Defining the Principles of PCR
PCR is a revolutionary method developed by Kary Mullis in the 1980s that relies on the natural ability of DNA polymerase to synthesize new strands of DNA complementary to a template strand.1 The power of PCR, however, lies in its capacity for specificity.
Since DNA polymerase can only attach a nucleotide onto a preexisting 3'-OH group, it absolutely requires a small, pre-designed oligonucleotide, known as a primer, to initiate synthesis.1
By designing two specific primers that flank a target region, researchers can ensure that only the desired sequence is amplified, accumulating billions of copies (amplicons) over typically 20 to 40 thermal cycles.1
Query: How does the initial discovery of a heat-stable polymerase affect the workflow compared to earlier replication techniques?
The invention of heat-stable polymerases, such as the widely adopted Taq polymerase, coupled with the development of automated thermal cyclers in the 1990s, fundamentally improved the efficiency and reliability of PCR.2 Previously, fresh enzyme had to be added after every high-temperature denaturation step. Today, the entire process is automated, requiring only a single, precise PCR Setup involving the necessary PCR reagents and robust PCR enzymes.3
1.2. The Three Steps of the PCR Cycling Parameters
A standard PCR is executed inside a thermal cycler - a device utilizing Peltier blocks to rapidly shift temperatures.4 Each cycle consists of three fundamental temperature steps 2:
1.2.1. Denaturation
The reaction mixture is heated to a high temperature, typically between 94°C and 98°C.5 This critical step breaks the weak hydrogen bonds holding the double-stranded DNA template together, separating it into two single strands that can then serve as templates for new synthesis.6 The initial denaturation step is often longer (e.g., 5 minutes) to ensure complete template unwinding and activation of certain Hot Start PCR enzymes.5
1.2.2. Primer Annealing
The temperature is rapidly lowered (often between 50°C and 65°C) to allow the forward and reverse primers to bind specifically to their complementary sequences on the single-stranded template.2 This is the step where specificity is determined, making the annealing temperature a crucial element of the PCR Cycling Parameters.
1.2.3. Extension (Elongation)
The temperature is raised again, usually to the optimal activity temperature of the polymerase (e.g., 72°C for Taq polymerase).5 The PCR enzymes then utilize the dNTP building blocks to synthesize a new DNA strand, starting from the 3’ hydroxyl group of the annealed primer, completing the cycle.1
Section 2: Building the Perfect Cocktail: A Deep Dive into PCR Reagents and Setup
Successful amplification hinges on the precise assembly of the reaction mixture. The PCR setup involves a delicate balance of concentration, purity, and volume (typically 50 μL) for all PCR reagents.5
2.1. Standard Components and Preparation
Experts often follow a standardized order of addition to mitigate issues like premature enzyme activity or non-specific binding: water, buffer, dNTPs, MgCl₂, template, primers, and finally, the Taq polymerase.5 Proper mixing and brief centrifugation are recommended to ensure all components are collected at the bottom of the tube.9
2.1.1. Primers and Template DNA
The DNA template must be high-quality and purified.8 While plasmid or viral templates require only 1 pg to 10ng, genomic templates typically require 1 ng to 1 μg.8 Excess DNA template is often a greater cause of non-specificity than too little, as it increases the likelihood of extraneous bands, particularly with high cycle numbers.8
Primers are the specificity drivers and should generally be 20 to 30 nucleotides long, ideally with 40% - 60% GC content.8 Crucially, the primer pair must have matching melting temperatures (𝑇𝑚) within 5°C to ensure they anneal simultaneously under the same conditions.8 Final primer concentration is typically 0.1–0.5 μM.5
2.1.2. dNTPs and PCR Buffer
Deoxynucleotide triphosphates (dNTPs) are the molecular building blocks, maintained at a final concentration of approximately 200 μM for each nucleotide.5 The PCR buffer is essential for stabilizing the reaction, maintaining optimal pH and ionic strength necessary for enzyme functionality.11
2.2. The Critical Regulatory Role of Magnesium Chloride
Magnesium Chloride is perhaps the most critical component in the PCR reagents mix, acting as an indispensable cofactor for DNA polymerase activity. The concentration of the magnesium ion, typically maintained between 0.1 mM and 0.5mM 5, directly dictates reaction output.
Query: How does magnesium ion concentration simultaneously impact enzyme activity and binding specificity?
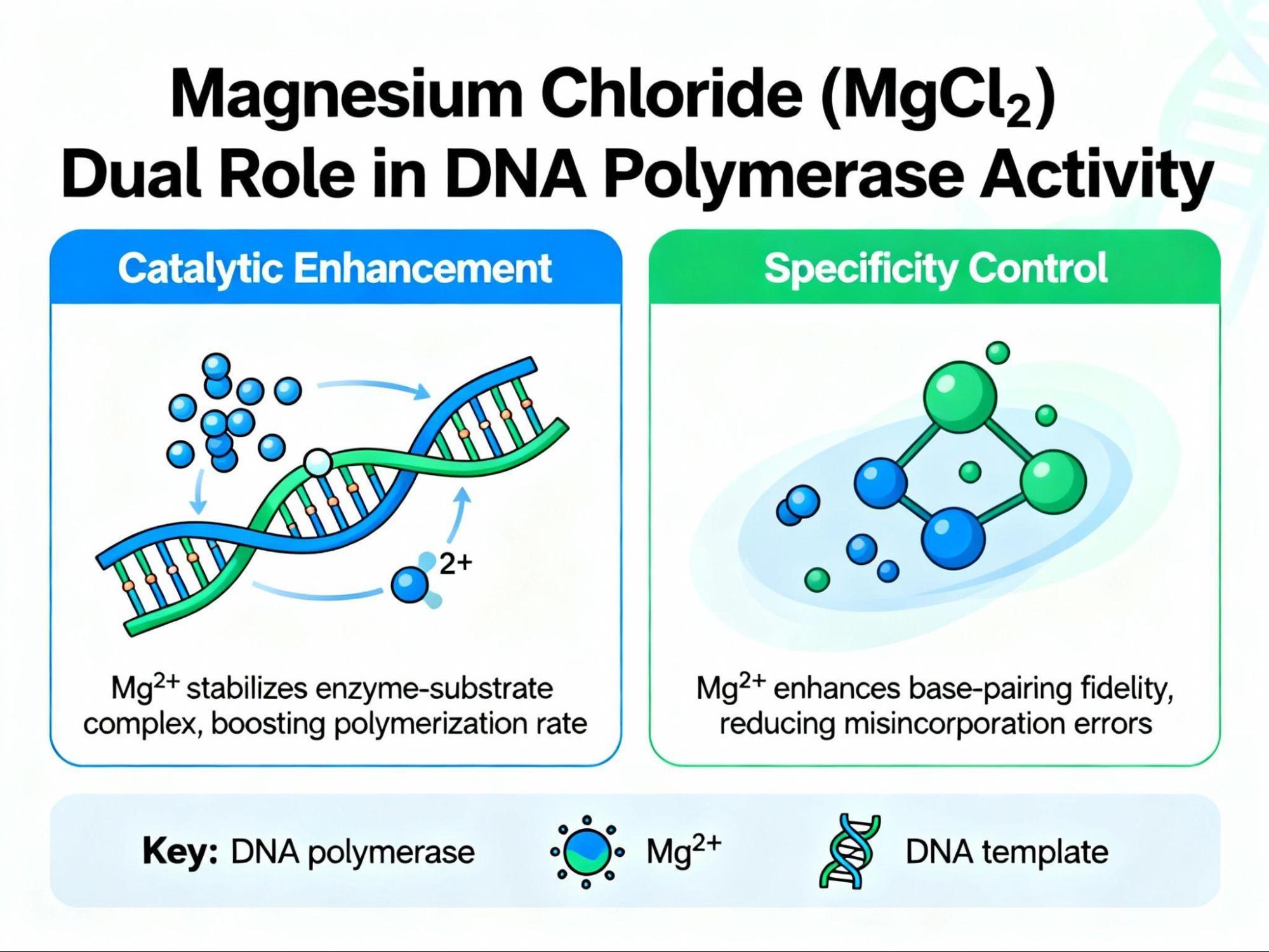
The magnesium ion plays a dual role:
- Catalytic Enhancement: This particular ion promotes the catalytic activity of the Taq DNA polymerase, increasing the overall amplification rate.12 Mechanistically, magnesium ion binds to the alpha-phosphate group of the dNTP, facilitating the necessary cleavage of the beta and gamma phosphates required for the formation of the phosphodiester bond with the 3’OH of the existing strand.13
- Specificity Control: Magnesium ion binds to the negatively charged phosphate groups in the DNA backbone. This binding reduces electrostatic repulsion between the two DNA strands, which, in turn, increases the 𝑇𝑚 and stabilizes the primer-template interaction.12
However, this dual function creates a delicate optimization challenge. While higher MgCl₂ concentrations increase productivity, they decrease specificity, often leading to the non-specific binding of primers and resulting in undesirable band smears on the gel.13
Conversely, too little MgCl₂ will result in poor amplification or complete failure due to insufficient enzyme catalysis or failure of proper primer annealing.13 Furthermore, if the DNA extract contains inhibitors, the MgCl₂ concentration might need to be increased to compensate for inhibitors that may sequester magnesium ions.13
| Component | Typical Final Concentration | Key Function |
|---|---|---|
| PCR Buffer (10x) | 1x | pH and ionic stability |
| MgCl₂ | 0.1 - 0.5mM (Optimized) | Cofactor for polymerase; specificity regulator 5 |
| dNTP Mix | 200 μM (each) | DNA building blocks 9 |
| Primers (F & R) | 0.1 - 0.5uM | Defines target start and end 8 |
| Genomic DNA Template | 1ng - 1ug | Target nucleic acid 8 |
| DNA Polymerase | 0.05/μL | Catalytic engine 9 |
Table 1: Essential PCR Reagent Concentrations for Standard 50 μL Reaction
Section 3: The Engine of Amplification: Selecting the Right PCR Enzymes
The selection of PCR enzymes has profound implications for the efficiency, speed, and, most importantly, the fidelity of the amplification process.
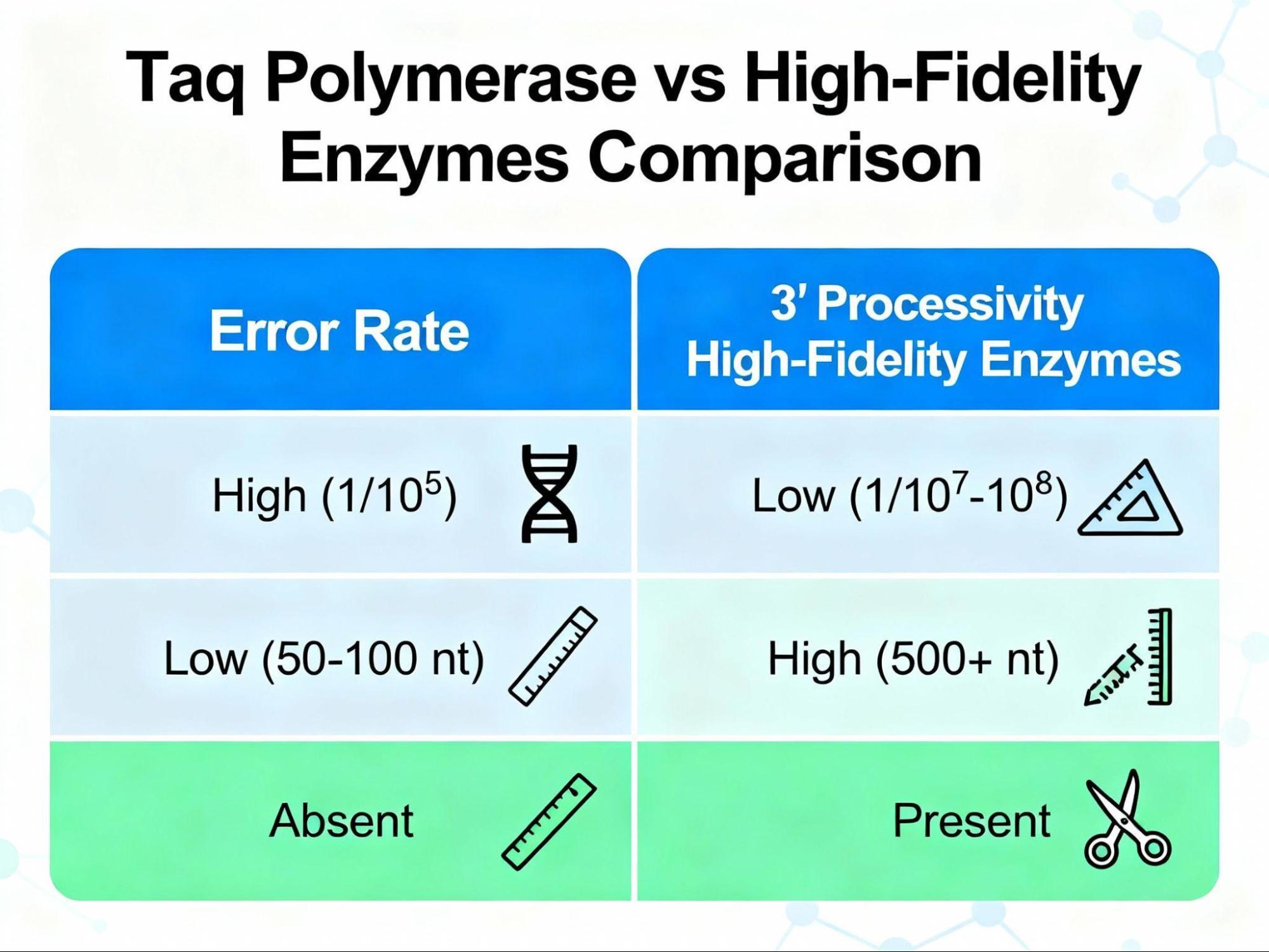
3.1. Taq Polymerase and the Fidelity Trade-Off
Taq DNA polymerase remains the industry standard for routine work due to its robust thermostability, which allows it to remain intact and active throughout the repeated high-temperature cycles.2
However, Taq lacks (3’ → 5’) exonuclease proofreading activity. Without this correction mechanism, Taq exhibits a relatively high base substitution rate, meaning it is prone to incorporating errors during synthesis. While suitable for simple screening and diagnostic applications, this error rate makes it unsuitable for sensitive downstream applications like cloning or sequencing, where the integrity of the generated sequence is paramount. 11
3.2. High-Fidelity Enzymes: Precision in Molecular Biology
For applications demanding high precision, specialized PCR enzymes with proofreading capability are essential. High-fidelity polymerases, often derived from Archaea (such as P fu or variants like Q5), possess an intrinsic 3’ → 5’ exonuclease domain.15 This domain functions as a quality control mechanism, allowing the enzyme to detect and excise misincorporated nucleotides before continuing synthesis.
The impact of this proofreading is significant; for example, the presence of the 3’ → 5’ exonuclease domain was shown to provide a 125-fold decrease in error rate compared to its exonuclease-deficient counterpart.15 Top-tier enzymes, like Q5 High-Fidelity DNA Polymerase, have been demonstrated to be up to 280-fold more faithful than standard Taq.15
For researchers performing cloning or sequencing, selecting high-fidelity PCR enzymes is crucial for ensuring the scientific validity of the findings.11
Query: Does selecting a high-fidelity enzyme automatically guarantee a perfect PCR run?
Not necessarily. While high-fidelity polymerases ensure accuracy, they sometimes come with a trade-off in processivity. Many archaeal proofreading polymerases exhibit limited processivity (synthesizing fewer than 20 bases before dissociating) compared to Taq, which can require optimization – particularly, the use of longer extension times for larger amplicons.16
3.3. Hot Start Technology: Boosting Specificity from the Start
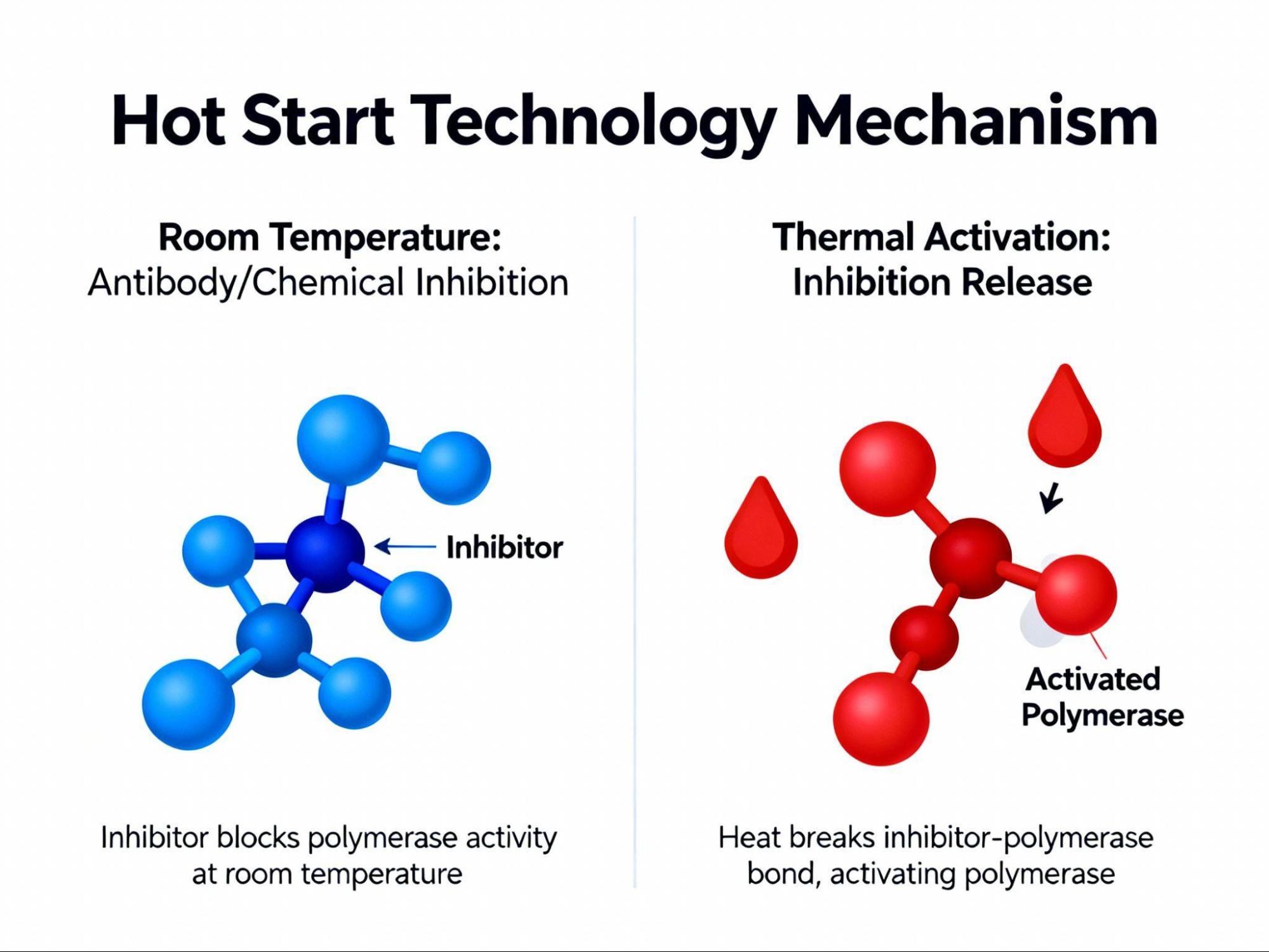
A significant challenge in PCR Setup is non-specific product formation, such as primer dimers, which can occur when the enzyme is active at lower temperatures during reaction assembly. Hot Start technology directly addresses this issue by inhibiting the polymerase at ambient temperatures.17
Hot Start PCR enzymes are typically bound by a chemical moiety, an antibody, or an aptamer that prevents activity until the enzyme is thermally activated during the initial high-temperature denaturation step.18 This strategy prevents the extension of primers that bind with low homology (mispriming) and prevents primer dimer formation, dramatically increasing the yield and specificity of the desired target fragment.17 This makes Hot Start technology invaluable for complex or high-throughput workflows.18
Section 4: Fine-Tuning the Machine: Optimizing PCR Cycling Parameters
Once the optimal PCR reagents and PCR enzymes are selected, achieving maximum yield and specificity requires careful optimization of the PCR cycling parameters.
4.1. Managing Denaturation Time
While the initial denaturation must be long enough to completely separate complex templates and activate hot start enzymes (up to 5 minutes) 6, the denaturation step in subsequent cycles must be minimized (typically 15-30 seconds at 95°C).5 Prolonged incubation at these high temperatures can lead to the loss of activity of certain enzymes, particularly standard Taq polymerase.6
4.2. Precision in Annealing Temperature (𝑇𝑎)
The annealing temperature is the primary lever for specificity. If non-specific bands are present, the PCR troubleshooting guide often suggests increasing the 𝑇𝑎.
- Optimal Range: The 𝑇𝑎 is typically set 5°C below the lowest primer's calculated 𝑇𝑚.
- Optimization: If spurious amplification occurs, researchers should test higher annealing temperatures. Increasing the 𝑇𝑎 enforces stricter binding requirements, destabilizing mismatches, and ensuring only perfectly complementary primers successfully anneal to the template.8 Running a temperature gradient on the thermal cycler is the most efficient way to pinpoint this optimal condition.20
New advancements in buffer chemistry utilize isostabilizing components that enhance the stability of the primer-template duplex, effectively allowing a universal annealing temperature (e.g., 60°C) even for primers with different 𝑇𝑚 values, thereby simplifying the PCR setup.6
4.3. Determining Extension Time
The extension step is performed at the enzyme's optimal temperature, often 68°C or 72°C.8 The duration of this step is directly proportional to the size of the target amplicon.
- The 1 kb Rule: A standard guideline is to use one minute of extension time per 1,000 base pairs (kb) of product.21 Products under 1kb usually require 45–60 seconds.8
- Necessity for Length: For long-range PCR (products greater than 3kb) or reactions involving many cycles (>30), longer extensions are required to ensure the enzyme completes synthesis on all templates.8 Insufficient extension time can result in incomplete products, which often appear as smears below the target band on a gel.6 For cloning applications using TA vectors, a long final extension step (up to 30 minutes) is often recommended to ensure proper 3’-dA tailing.6
4.4. Cycle Count
The total number of cycles used balances the desired yield against the risk of accumulating errors and non-specific products. The standard range is 20 - 35 cycles.22
- High Template Concentration: Use fewer cycles (25 - 30) when the template concentration is high.19
- Low Template Concentration: Use more cycles (30 - 35) when the template concentration is low.19
Excessive cycling increases the chance for non-specific products to be amplified exponentially in the later stages of the reaction.19
Section 5: Diverse Applications and Methods: Putting PCR to Work
The technological advances in PCR reagents and instrumentation have broadened the spectrum of PCR applications, making it a staple across life sciences and molecular diagnostics.23
5.1. Foundational PCR Applications
Routine endpoint PCR is foundational for numerous research tasks, including:
- Genotyping: Identifying genetic markers or mutations.24
- Cloning and Mutagenesis: Preparing DNA fragments for vector insertion or generating targeted changes (often requiring high-fidelity PCR enzymes). Researchers interested in accurate protein expression can browse our selection of highly validated PCR kits.
- Infectious Disease Diagnosis: Early applications included detection of pathogens like Mycobacterium tuberculosis and Chlamydia trachomatis.25
5.2. Specialized PCR Methods for Enhanced Performance
When dealing with highly sensitive targets or challenging templates, specialized PCR methods are employed to boost specificity and sensitivity.
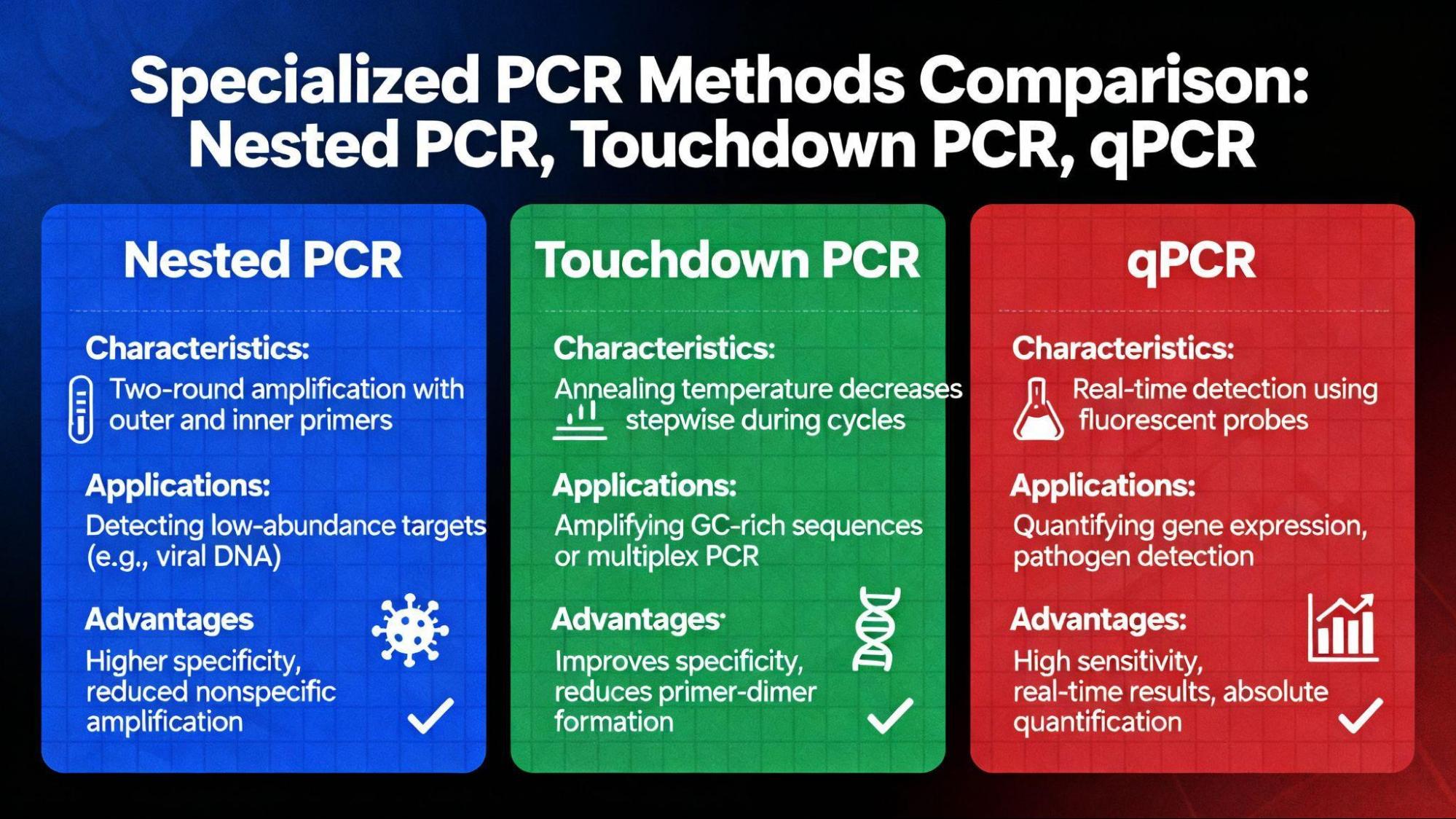
5.2.1. Nested PCR
Nested PCR involves two sequential PCR runs using two pairs of primers. The second set of (inner) primers binds within the product generated by the first set of (outer) primers.26 This method significantly increases specificity by requiring two rounds of successful primer binding and is highly sensitive for low-copy number detection.27
5.2.2. Touchdown PCR (TD-PCR)
TD-PCR is a powerful technique for optimizing specificity, especially in cases where primer design is difficult. The method involves starting the high 𝑇𝑎 (e.g., 65°C) and progressively lowering it (e.g., 0.5°C every cycle) until a desired 𝑇𝑎 minimum is reached, followed by several cycles at that final low temperature.26 The initial high-stringency cycles ensure that only highly specific primer binding occurs, enriching the correct product, which is then amplified in the final, lower-temperature cycles.
5.3. Real-Time Quantitative PCR (qPCR)
Quantitative PCR (qPCR) fundamentally changed how nucleic acids are analyzed by allowing quantification rather than just detection.28
- Mechanism of Quantification: Unlike endpoint assays, which measure the product accumulated after a fixed number of cycles, qPCR monitors fluorescence reporters (dyes or probes) in real-time.28 The cycle number at which the fluorescence signal crosses a detectable threshold (the Cq value) is inversely proportional to the initial quantity of the target DNA or RNA.28
- Superior Sensitivity: Clinical data indicate that RT-qPCR offers higher sensitivity and speed compared to Nested-PCR for applications such as diagnosing acute leukemias, requiring a smaller amount of PCR reagents and less time.27 This method is crucial for diagnostics and disease monitoring, allowing quick and reliable assessment.27
qPCR kits utilizing probe-based or dye-based chemistries are available, often falling under our category of “other kits“, which serve complementary research needs.
Section 6: Workflow Efficiency and Quality: Choosing the Best PCR Kits
The increasing demand for reliable and reproducible molecular analysis, particularly in clinical and high-throughput settings, has accelerated the adoption of commercial PCR kits and master mixes.30
6.1. Advantages of Pre-Optimized PCR Kits and Master Mixes
Commercial master mixes typically contain pre-optimized concentrations of buffer, dNTPs, MgCl₂, and DNA polymerase (often hot start versions).18
- Minimizing Variables: Ready-to-use master mixes are essential for high-throughput or time-sensitive workflows, as they drastically reduce pipetting errors and save preparation time.11 By providing consistent, lot-to-lot performance, they improve reproducibility across replicates, a core requirement for robust diagnostics and publication-quality research.11
- Market Dynamics: The kits and reagents sector dominated the molecular biology market in 2024, accounting for 65.52% of the revenue share, highlighting the industry preference for standardized, convenient solutions.30
6.2. The Role of Reagent Purity and Inhibitor Tolerance
The reliability of quantitative assays depends on the efficiency and precision of the amplification, which, in turn, is determined by the quality of the PCR reagents.31
- Quality Control: High-quality reagents are manufactured under stringent quality control standards to ensure the stability of stabilizers (like BSA) and the consistency of crucial ion concentrations.11 When comparing products, researchers must check for optimized buffer compositions and appropriate magnesium levels.11
- Inhibitor Tolerance: For researchers working with crude or complex biological samples (e.g., soil, blood, or tissue lysates), choosing PCR enzymes and master mixes that are formulated to tolerate common PCR inhibitors is vital.11 Reagents with enhanced inhibitor tolerance help maintain high assay sensitivity and specificity despite complex sample matrices.
At AAA Biotech, we prioritize stringent quality control across all of our products, ensuring optimal performance across diverse applications.
Section 7: The Practical Lab Fix: Your PCR Troubleshooting Guide
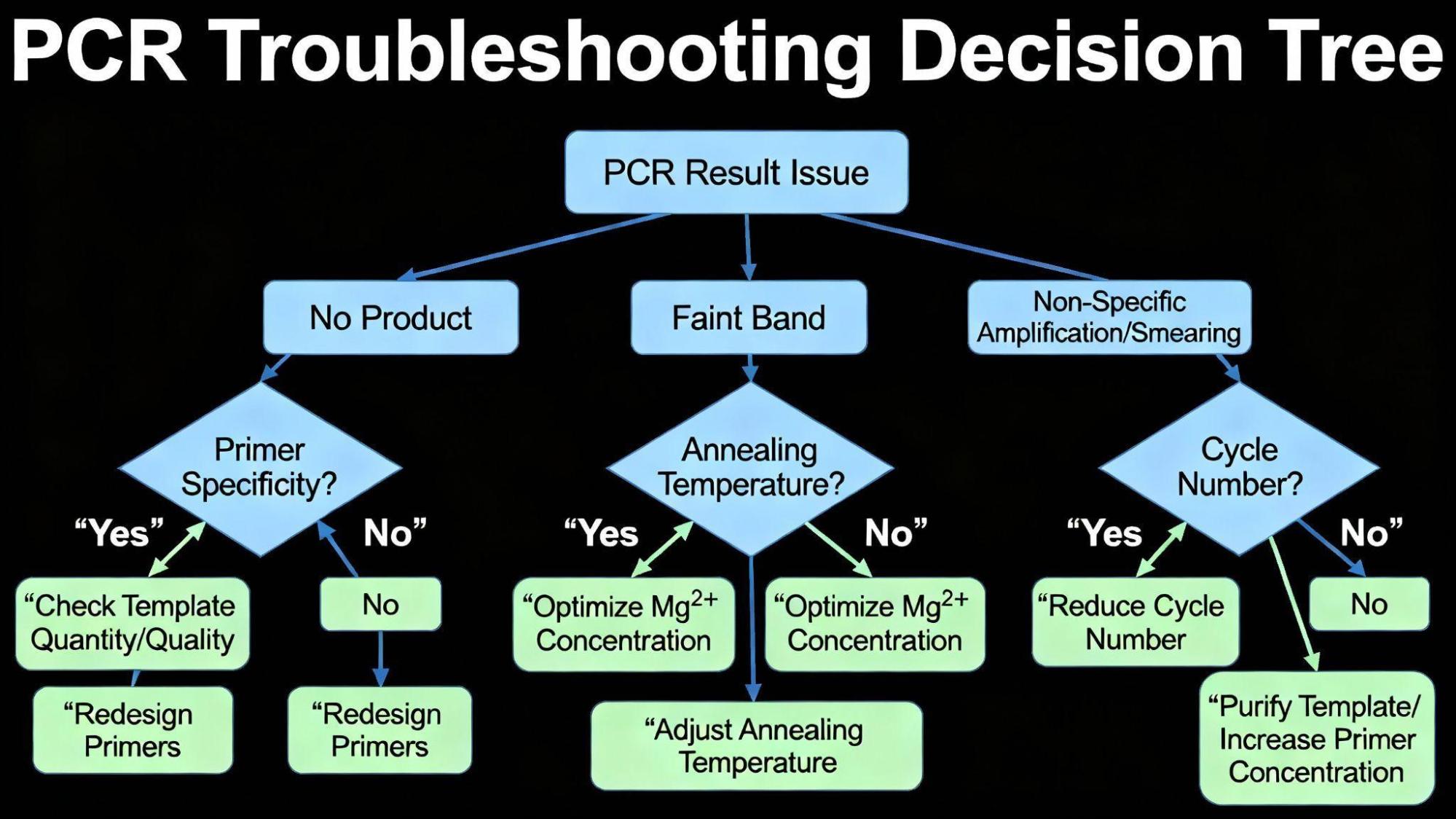
Even with optimal reagents, deviations from expected results are common. Effective PCR troubleshooting guide strategies involve systematically adjusting the most critical variables: annealing temperature, MgCl₂ concentration, and cycle number.
7.1. Scenario A: No Product or Faint Band
“Failure to amplify” typically points to problems in template accessibility, primer binding, or enzyme function.
7.1.1. Addressing Template and Cycle Counts
If the band is faint or absent, the concentration of the DNA template may be too low, or the DNA may be degraded.10 If the quality is acceptable, the solution is often to increase the template input (up to 1 μg for genomic DNA) or increase the number of amplification cycles.10
7.1.2. Optimizing 𝑇𝑎 and MgCl₂
If “too few cycles” is not the issue, consider the temperature settings. If the annealing temperature is too high, it prevents the primers from binding to the template, leading to amplification failure.20 A systematic annealing temperature gradient should be used to find the optimal 𝑇𝑎.20 Likewise, insufficient MgCl₂ severely limits the catalytic activity of the polymerase, resulting in weak or no product.13 A small increase in MgCl₂ may restore activity.13
7.2. Scenario B: Non-Specific Amplification or Smearing
The appearance of multiple bands or smearing indicates a loss of specificity, meaning the primers are binding to non-target regions and being amplified.
7.2.1. Increasing Stringency
Non-specific binding frequently occurs because the annealing temperature is too low, allowing primers to bind promiscuously.19 The primary solution in this PCR troubleshooting guide is to raise the 𝑇𝑎, which increases the stringency of the reaction.19
7.2.2. Adjusting Reagent Concentrations and Cycle Count
Excessive template concentration is a common cause of non-specific binding and should be reduced.10 Similarly, excessive MgCl₂ concentration drastically reduces specificity, leading to non-specific binding, primer dimers, and extra bands.13
Reducing MgCl₂ concentration can restore specificity.13 Furthermore, excessive cycling (above 35 cycles) increases the opportunity for non-specific products, which initially exist at very low concentrations, to be amplified to detectable levels.22 Reducing the cycle count is often recommended for cleaner results.19 Finally, if primer dimers are forming during reaction setup, employing a Hot Start enzyme technology is the best preventive measure.17
| Observed Problem | Primary Cause | Optimization Strategy |
|---|---|---|
| No Band/Faint Product | Template quantity too low or degraded | Increase template/cycles; re-extract DNA 10 |
| No Band/Faint Product | Annealing temperature too high or MgCl₂ too low | Lower annealing temperature (run gradient); increase MgCl₂ slightly 13 |
| Smearing/Multiple Bands | Annealing temperature too low or MgCl₂ too high | Raise annealing temperature; reduce MgCl₂ concentration 13 |
| Smearing/Multiple Bands | Excessive Template or Cycles | Reduce template quantity; limit cycles to 25–3510 |
Table 2: Key Troubleshooting Solutions for Standard PCR Problems
Conclusion
Mastering PCR is a journey that moves from understanding the fundamental principles of PCR to executing a meticulous PCR setup and skillfully optimizing its cycling parameters. The success of any molecular assay, whether standard screening or advanced quantitative diagnostics, is inherently tied to the quality and performance of the chosen PCR reagents and enzymes.
Precision in controlling MgCl₂ concentration, selecting high-fidelity polymerases when sequence integrity is vital, and utilizing modern PCR kits (like master mixes with Hot Start capability) are the hallmarks of a successful laboratory workflow.
For researchers seeking reliability and excellence across all PCR applications and PCR methods, AAA Biotech offers a comprehensive portfolio of high-purity reagents and pre-optimized PCR kits designed to minimize errors and ensure accurate, reproducible results.
Ready to move beyond troubleshooting and amplify your success? Contact us today to discuss your specific requirements.
Faq's
What is the main difference between Taq and high-fidelity PCR Enzymes?
High-fidelity enzymes possess 3’ → 5’ exonuclease proofreading activity, resulting in significantly lower error rates (up to 280X better than Taq), which is essential for cloning applications.
Why must I run a negative control during PCR setup?
A negative control (containing all reagents but no template DNA) is essential to detect reagent contamination, thereby preventing false positive results that could invalidate the experiment.
How does increasing MgCl₂ concentration affect the PCR reaction?
Increased MgCl₂ boosts DNA polymerase activity but simultaneously decreases primer specificity, which often leads to non-specific binding and the appearance of undesired bands.
What is the key advantage of using a commercial PCR Kit master mix?
Master mixes reduce the number of pipetting steps necessary, thereby minimizing human error, saving time, and greatly improving the reproducibility and consistency of results across replicates.
What should I do first if my PCR result shows non-specific bands?
The first step should be to increase the annealing temperature slightly to boost stringency, and ensure that the reaction is not running for an excessive number of cycles (typically limited to 30–35).
Works Cited/ References
- Polymerase Chain Reaction (PCR) - NCBI - NIH
- Polymerase chain reaction - Wikipedia
- PCR Technology: Key Milestones in Development and Maturation | Thermo Fisher Scientific
- PCR Thermal Cyclers Education | Thermo Fisher Scientific - US
- PCR Protocol - PCR Steps - PCR Design Tool | GenScript
- PCR Cycling Parameters—Six Key Considerations for Success | Thermo Fisher Scientific
- Performance Evaluation of Thermal Cyclers for PCR in a Rapid Cycling Condition
- Guidelines for PCR Optimization with Taq DNA Polymerase - NEB
- Standard PCR Protocol - Sigma-Aldrich
- Weak PCR Band Results or Smearing - What To Do - GoldBio
- PCR Reagent Selection 101: Expert Tips for Accuracy and Efficiency - Pro Lab Supply Corp
- What is the role of MgCl2 in PCR? - ResearchGate
- What Is the Role of MgCl2 in PCR Amplification Reactions? - Excedr
- The role of MgCl2 in PCR - simple animated HD
- Polymerase Fidelity: What is it, and what does it mean for your PCR? - NEB
- High-Fidelity PCR Enzymes: Properties and Error Rate Determinations - Agilent
- How is Hot-Start Technology Beneficial For Your PCR
- Hot Start Taq 2X Master Mix - NEB
- Troubleshooting Non-Specific Amplification - Bento Lab
- No Bands - Genotyping | The Jackson Laboratory
- Optimization of Annealing Temperature and other PCR Parameters - Genaxxon bioscience
- PCR Troubleshooting | Bio-Rad
- PCR: The Technology That Launched Biotechnology - NSF Impacts
- PCR Tests: MedlinePlus Medical Test
- PCR-based diagnostics for infectious diseases: uses, limitations, and future applications in acute-care settings - PMC - PubMed Central
- Comparison of standard PCR with hemi-nested touchdown (TD) PCR. (A)... | Download Scientific Diagram - ResearchGate
- Nested-PCR vs. RT-qPCR: A Sensitivity Comparison in the Detection of Genetic Alterations in Patients with Acute Leukemias - MDPI
- Essentials of Real-Time PCR | Thermo Fisher Scientific - US
- Real-Time Polymerase Chain Reaction: Current Techniques, Applications, and Role in COVID-19 Diagnosis - PubMed Central
- Molecular Biology Enzymes, Kits & Reagents Market Size & Share Analysis, 2030
- Validating Real-Time Polymerase Chain Reaction (PCR) Assays - PMC - PubMed Central

 Cynthia
Cynthia