In this Article
- Which Bacteriophages Are Used in Phage Display?
- Types of Phage Display Antibody Libraries
- Phage Display Workflow: Step-by-Step
- Phage Display vs. Recombinant Antibody: Understanding the Relationship
- Phage Display Monoclonal Antibodies: From Hit to IND
- Applications of Phage Display Technology
- Phage Display Methods and Protocols: Technical Considerations
- Advantages and Limitations of Phage Display for Antibody Generation
- Phage Display vs. Other Antibody Library Discovery Platforms
- Phage Display Antibody Discovery Against Infectious Diseases and Toxins
- Next-Generation Advances in Phage Display Antibody Technology
- Conclusion
All of the products listed in AAA Biotech’s catalog are strictly for research-use only (RUO).

Quick Summary
Phage display is an in vitro antibody discovery platform that fuses antibody fragments to bacteriophage coat proteins, enabling iterative selection of high-affinity binders from libraries exceeding 1010 variants. It bypasses animal immunization, produces fully human recombinant antibodies, and underpins the discovery of blockbuster therapeutics, including adalimumab, the world's best-selling monoclonal antibody.
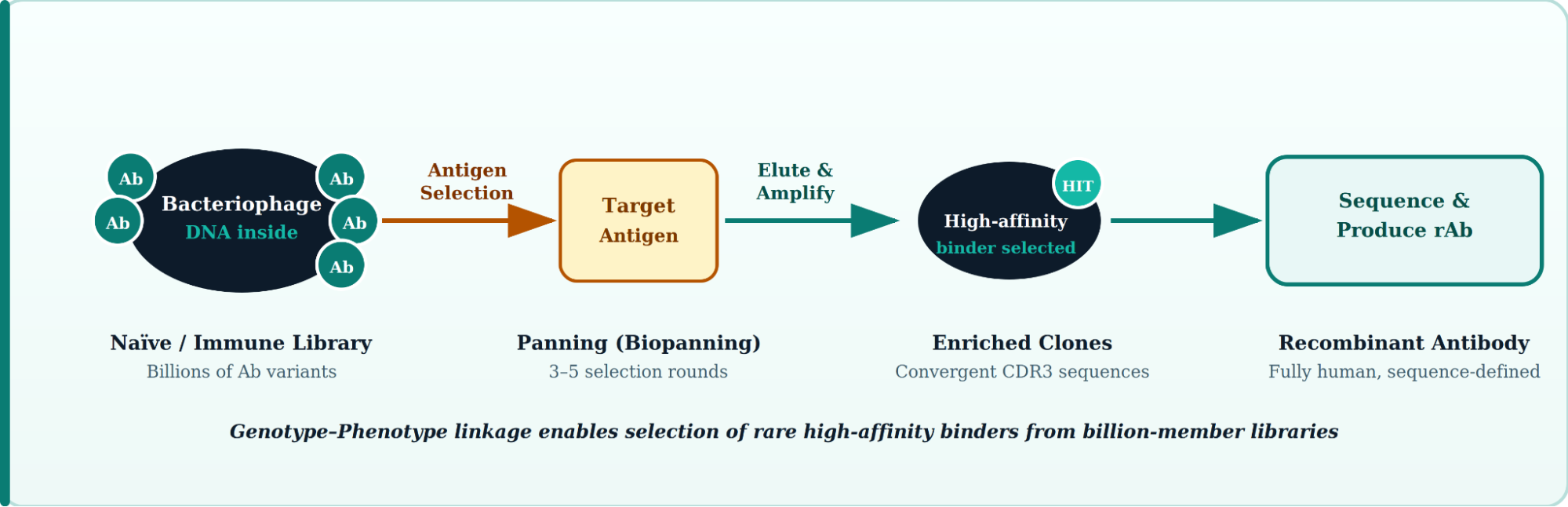
Phage display is a molecular selection technology in which a peptide or antibody fragment is genetically fused to the coat protein of a bacteriophage, physically linking each antibody's genetic code to its binding function on the same viral particle. This genotype–phenotype linkage allows researchers to screen enormous combinatorial libraries often containing more than one billion distinct clones against virtually any antigen, selecting rare, high-affinity binders with precision that traditional immunization cannot match.
First described by George P. Smith in 1985 and later expanded into the antibody field by John McCafferty and Greg Winter's laboratories in the early 1990s, phage display antibody technology earned its pioneers the 2018 Nobel Prize in Chemistry.
Today, phage display antibody discovery has matured into the dominant in vitro platform for producing fully human, sequence-defined, reproducible recombinant antibody assets critical to modern diagnostics, therapeutics, and research reagent development.
Key Definition: In phage display, the antibody's variable domains (scFv or Fab) are displayed on the phage surface while the encoding DNA resides inside, making each particle a self-contained molecular identity card that can be amplified, sequenced, and engineered.
Which Bacteriophages Are Used in Phage Display?
Not all phages are equal for antibody display. The choice of phage determines library capacity, display valency, and downstream workflow efficiency.
| Phage Type | Display Protein | Valency | Best For | Key Limitation |
|---|---|---|---|---|
| M13 filamentous | pIII (3–5 copies) or pVIII (2,700 copies) | Monovalent (pIII) / Polyvalent (pVIII) | Antibody fragment display (scFv, Fab); most widely used | Large inserts reduce infectivity via pVIII |
| T4 | Hoc / Soc proteins | High (up to 900 copies) | Vaccine development, multi-antigen display | Less established for antibody selection |
| T7 lytic | 10B coat protein | Variable | Targets requiring cytoplasmic folding | Not suitable for eukaryotic expression systems |
| Lambda (λ) | gpD tail protein | High | Peptide libraries, cDNA libraries | Lower yield; complex packaging |
In practice: Over 90% of phage display antibody workflows use M13 filamentous phage with the pIII coat protein, because monovalent display ensures that binders are selected on true affinity, not avidity.
Types of Phage Display Antibody Libraries
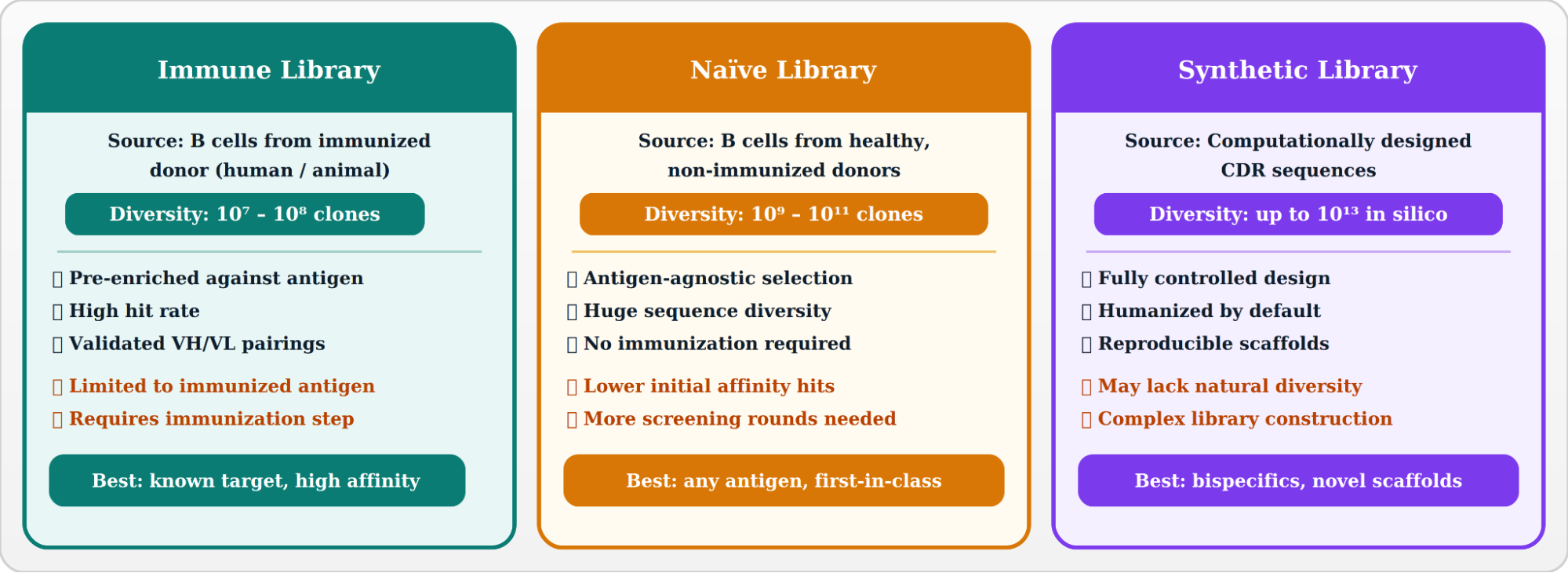
The depth and composition of the phage display antibody library dictate which antigens can be targeted and the quality of hits obtained. Three principal library classes exist:
Antibody Fragment Formats Displayed on Phage
- scFv (Single-Chain Variable Fragment): VH and VL domains connected by a flexible peptide linker (~15 aa Gly₄Ser repeat). Compact, easy to clone, the most common phage display format.
- Fab (Fragment Antigen-Binding): Full heavy-chain Fd + light chain. Closer to native antibody geometry; better structural validation for therapeutic development.
- VHH / Nanobodies: Single-domain antibodies from camelid heavy-chain-only antibodies. Exceptional stability and tissue penetration; increasingly used in phage display.
- dAb (Domain Antibodies): Smallest functional antibody unit; used for target engagement in difficult epitopes.
Phage Display Workflow: Step-by-Step
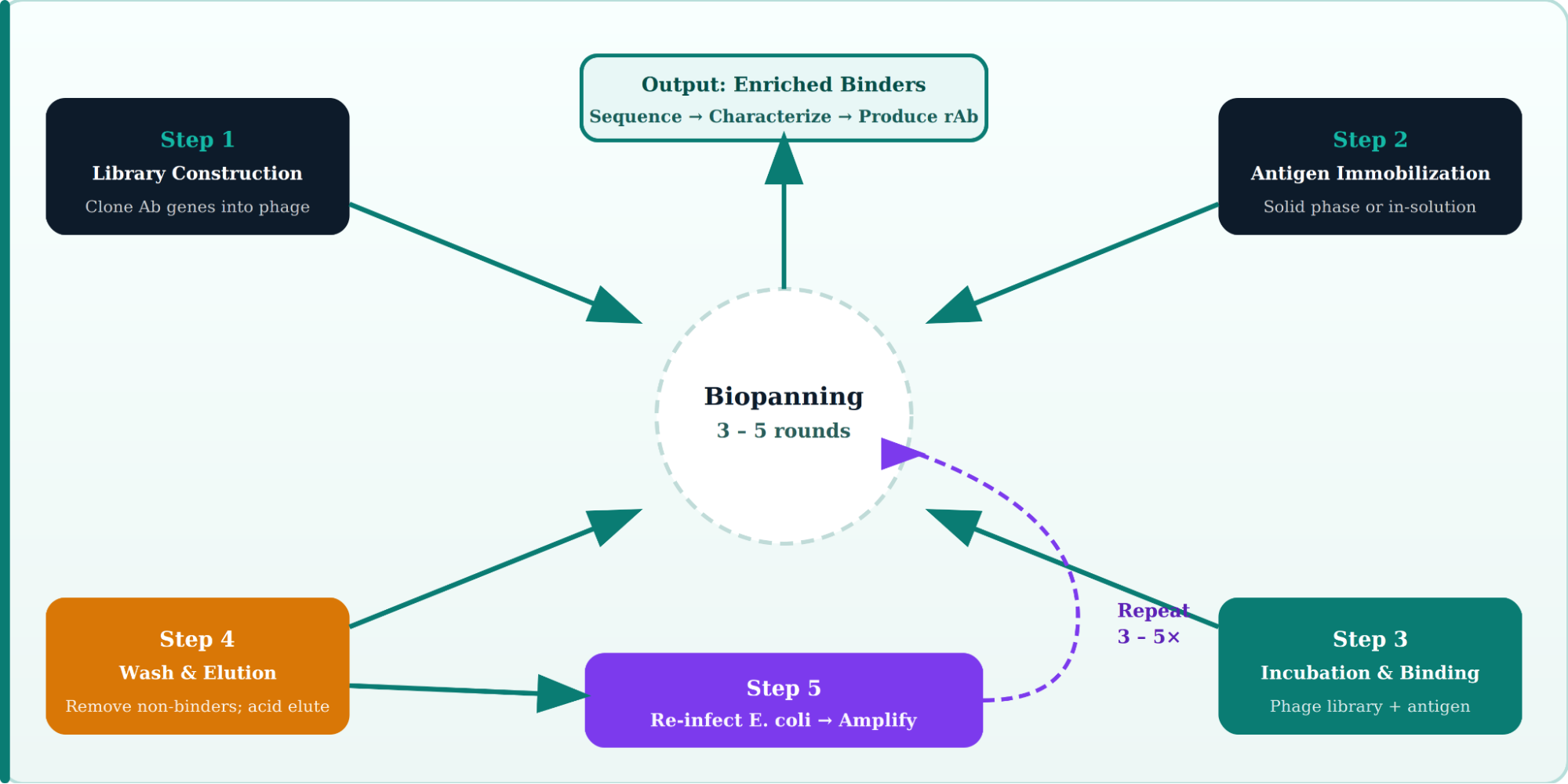
The phage display workflow follows a cyclical process called biopanning. Each round enriches binders by several orders of magnitude, typically requiring 3–5 selection rounds before confident hits are identified.
Detailed Steps in the Phage Display Antibody Production Pipeline
1. Library Construction
- RNA is extracted from B cells (peripheral blood lymphocytes, spleen, or bone marrow).
- cDNA is synthesized by reverse transcription, and VH/VL genes are amplified by RT-PCR using degenerate primers covering all human germline families.
- Antibody genes are cloned into phagemid vectors (e.g., pCANTAB, pHEN) in-frame with the pIII gene.
- E. coli is transformed, and libraries are packaged by co-infection with helper phage (M13KO7 or VCSM13).
2. Antigen Presentation Strategy
- Solid-phase panning: Antigen adsorbed onto immunotubes or magnetic beads. Simple; best for stable, purified proteins.
- Solution-phase panning: Biotinylated antigen + streptavidin beads. Better orientation control; preferred for membrane proteins.
- Cell-based panning: Live cells presenting native conformational epitopes. Critical for GPCR and other multi-pass membrane targets.
- Counter-selection (subtractive panning): Pre-incubate with off-target cells/proteins to deplete cross-reactive clones before antigen selection.
3. Washing and Elution
- Stringency is gradually increased across rounds (lower pH elution, higher salt, competitor antigen).
- Elution methods: low pH (glycine-HCl, pH 2.2), protease cleavage, antigen competition, or electroelution.
- Trypsin elution can be used when the pIII region contains a protease cleavage site.
4. Re-infection and Amplification
- The eluted phage infects exponential-phase E. coli TG1 or SS320 cells.
- Superinfection with helper phage packages phagemid DNA into new phage particles.
- Overnight culture yields ~10¹² phage/mL, ready for the next round.
5. Screening and Hit Identification
- Phage ELISA on individual colonies (96- or 384-well format) identifies antigen-positive clones.
- Positive clones are sequenced by Sanger or next-generation sequencing (NGS).
- Unique CDR3 sequences are clustered; diverse clones advanced to soluble antibody expression.
Phage Display vs. Recombinant Antibody: Understanding the Relationship
A common point of confusion is the distinction between phage display antibody and recombinant antibody. These are not opposing categories; they describe different aspects of the same process.
| Dimension | Phage Display Antibody |
Recombinant Antibody (Broadly) |
Traditional Hybridoma mAb |
|---|---|---|---|
| Discovery route | In vitro biopanning from phage libraries | Any in vitro molecular biology approach | In vivo immunization + cell fusion |
| Animal required? | No (naïve/synthetic) or optional (immune) | No | Yes |
| Humanization needed? | Not for fully human libraries | Depends on origin | Yes (murine → human) |
| Sequence defined? | Yes - from day 1 | Yes | Post-hoc sequencing required |
| Library diversity | Up to 10¹¹ variants | Varies by method | N/A |
| Epitope control | High (cell-based, competitive panning) | Variable | Low |
| Reproducibility | Excellent (sequence-defined) | Excellent | Poor (clonal drift) |
| Batch-to-batch consistency | High | High | Variable |
| Toxic/non-immunogenic targets | Accessible | Accessible | Difficult |
| Regulatory precedent | Strong (adalimumab, etc.) | Strong | Long history |
Bottom line: All phage-display-derived antibodies are recombinant antibodies by definition. However, not all recombinant antibodies originate from phage display; other platforms include yeast display, ribosome display, and mammalian cell display. Phage display remains the most validated and widely deployed of these antibody library discovery platforms.
Phage Display Monoclonal Antibodies: From Hit to IND
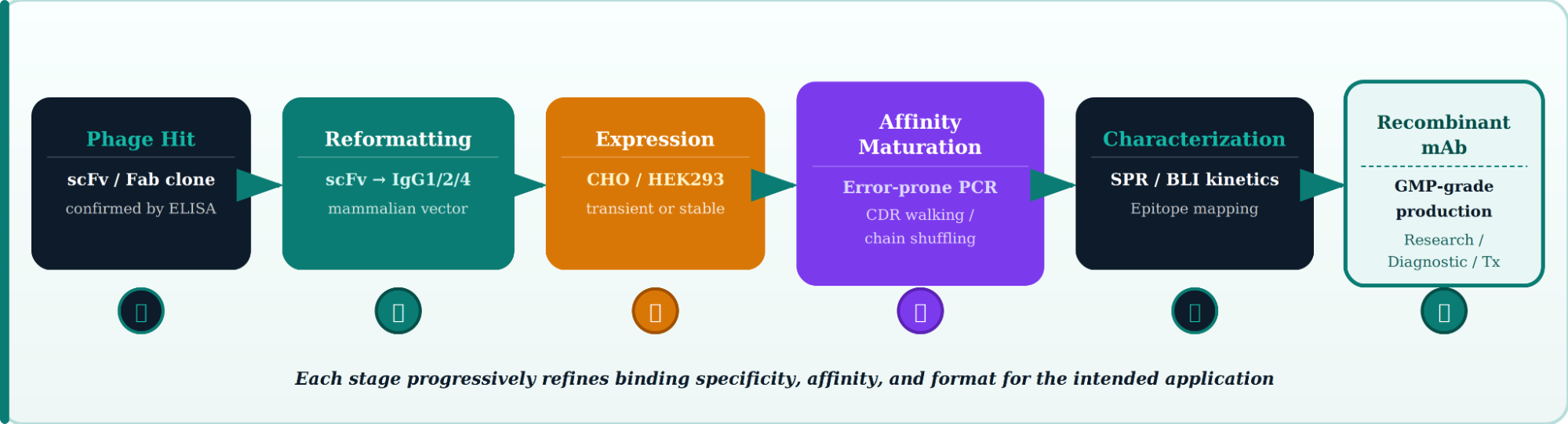
Once a phage display hit is confirmed by ELISA and sequencing, the journey to a clinical-grade phage display monoclonal antibody involves several precision engineering steps.
Affinity Maturation Strategies After Phage Display
- CDR walking mutagenesis: Systematic single-position mutations across CDR loops; identifies key contact residues and improves KD.
- Error-prone PCR (epPCR): Random mutations across the V-region; combined with additional rounds of phage display to select enhanced binders.
- Light/heavy chain shuffling: Original VH (or VL) is paired with a library of partner chains; exploits combinatorial diversity for improved affinity.
- Look-through mutagenesis (LTM): Systematic CDR residue substitution with a defined panel of 9 amino acids; rapid affinity optimization.
Applications of Phage Display Technology
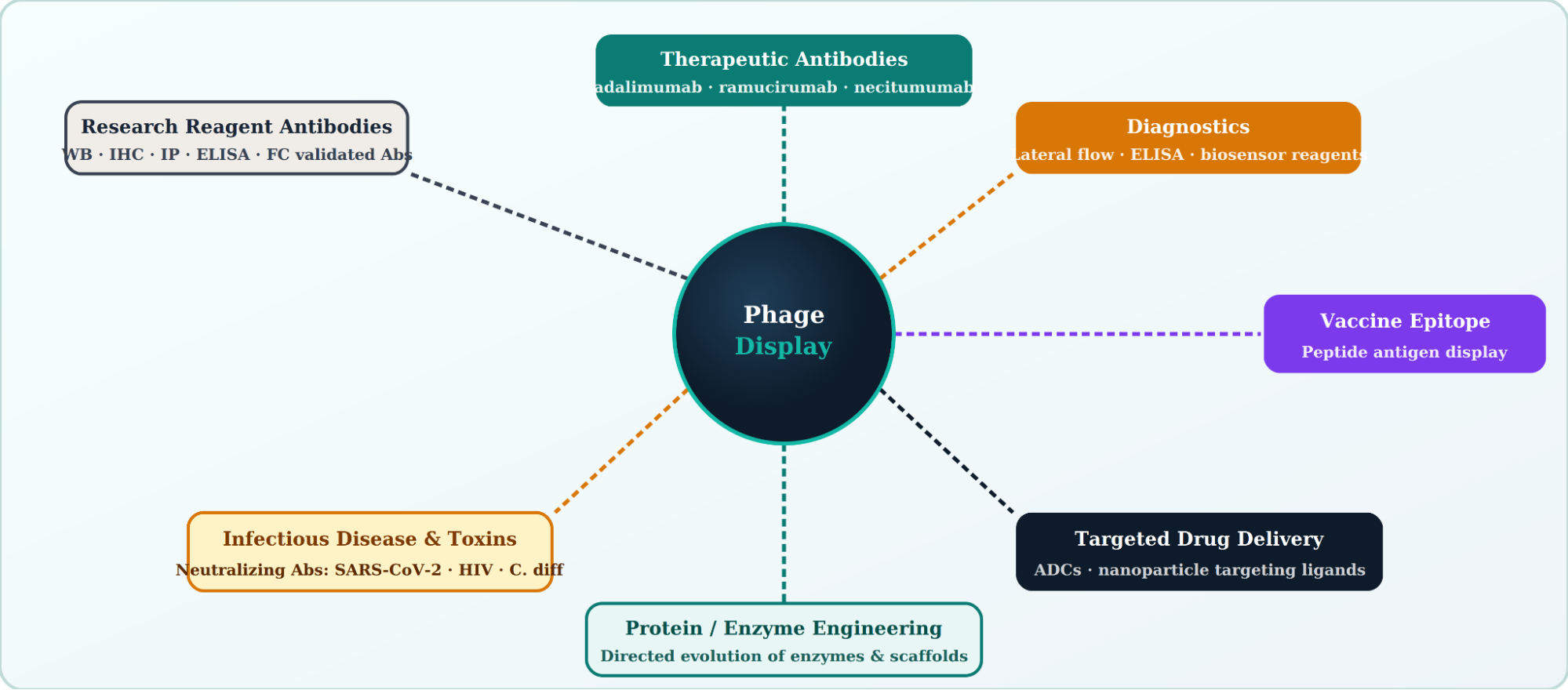
The versatility of phage display extends well beyond monoclonal antibody discovery. Its applications of phage display now span nearly every branch of modern biomedicine.
Approved Antibody Drugs Derived from Phage Display
| Drug Name | Target | Indication | Approval | Format |
|---|---|---|---|---|
| Adalimumab (Humira®) | TNF-α | RA, Crohn's, psoriasis | FDA 2002 | Fully human IgG1 |
| Ramucirumab (Cyramza®) | VEGFR-2 | Gastric, NSCLC, HCC | FDA 2014 | Fully human IgG1 |
| Necitumumab (Portrazza®) | EGFR | Squamous NSCLC | FDA 2015 | Fully human IgG1 |
| Olaratumab (Lartruvo®) | PDGFRα | Soft tissue sarcoma | FDA 2016 | Fully human IgG1 |
| Avelumab (Bavencio®) | PD-L1 | Merkel cell carcinoma, UC | FDA 2017 | Fully human IgG1 |
Phage Display Methods and Protocols: Technical Considerations
Library Quality Metrics
- Library size: A functional library for therapeutic discovery requires ≥10⁹ independent transformants. Smaller libraries (<10⁷) limit diversity and reduce hit rates.
- Insert frequency: ≥80% of clones should carry a correct in-frame insert. Libraries with higher frameshifts or stop codons waste selection capacity.
- Representation: Deep sequencing of naive libraries should confirm broad VH and Vκ/Vλ germline coverage, particularly of the CDR3 length distribution.
Panning Condition Optimization
- Antigen concentration: Start at 1-10 µg/mL for solid-phase panning; reduce by ~10-fold each round to increase stringency.
- Blocking agents: 2-4% MPBS (milk in PBS) or 1% BSA. Avoid casein for antibodies targeting milk-derived antigens.
- Wash stringency: Increase PBST (0.05% → 0.5% Tween-20) across rounds; add competitor protein in later rounds for specificity selection.
- pH elution parameters: 100 mM glycine-HCl pH 2.2 (10-min incubation), immediately neutralized with 1M Tris-HCl pH 8.0. Alternative: competitive elution with 100–500 µg/mL soluble antigen.
Screening Format Options
| Screening Method | Throughput | Sensitivity | Function |
|---|---|---|---|
| Phage ELISA (monoclonal) |
96–384 clones/plate | High | Antigen binding only |
| Soluble scFv ELISA | 96–384 clones/plate | High | Binding; early format switch |
| NGS (Illumina) of enriched pool | Millions of reads | Very high | Full diversity landscape |
| Protein microarray | Thousands of antigens | Medium | Specificity profiling |
| Cell-based FACS sorting | High (10⁶+ events) | High | Native conformation binding |
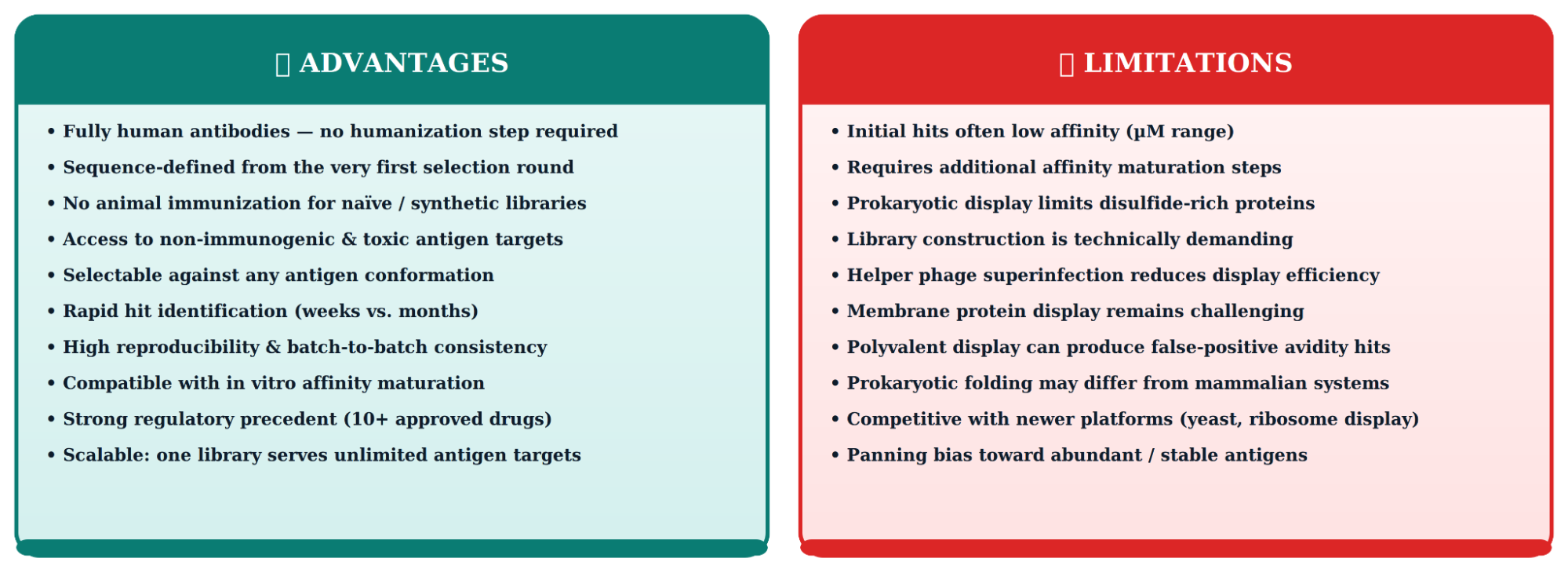
Advantages and Limitations of Phage Display for Antibody Generation
Phage Display vs. Other Antibody Library Discovery Platforms
| Platform | Display System |
Library Size |
Best Format | Key Advantage Over Phage |
Key Disadvantage vs. Phage |
|---|---|---|---|---|---|
| Phage Display |
M13, T7, λ phage | 10⁹–10¹¹ | scFv, Fab, VHH | — | — |
| Yeast Display |
S. cerevisiae Aga2p |
10⁷–10⁹ | scFv, Fab | FACS-based quantitative selection; eukaryotic folding | Smaller library; slower growth |
| Ribosome Display |
Ribosome-mRNA-protein complex | 10¹³–10¹⁵ | scFv, peptides | Largest diversity; fully cell-free | Less stable; no in vivo amplification |
| Mammalian Cell Display |
CHO, HEK293 surface |
10⁶–10⁸ | IgG (full-length) |
Native IgG format from start; glycosylation | Smallest library; expensive |
| mRNA Display |
Puromycin-linked mRNA | 10¹³ | Peptides, domains | Very large peptide libraries | Limited to small proteins/peptides |
Phage Display Antibody Discovery Against Infectious Diseases and Toxins
Research published in Frontiers in Cellular and Infection Microbiology (Roth et al., 2021) documented extensive application of phage display antibody generation to rapidly isolate neutralizing antibodies against emerging pathogens, where timelines are too compressed for traditional immunization-based approaches.
Notable Examples
- SARS-CoV-2: Phage display against the receptor-binding domain (RBD) and spike protein S1 yielded potent neutralizing scFv and nanobody clones within 4-6 weeks of antigen availability, critical during pandemic response.
- Clostridium difficile toxins A and B: Fully human neutralizing Fabs selected from naïve scFv libraries, bypassing the difficulty of producing active toxin for animal immunization.
- HIV-1 gp120/gp41: Phage display has identified broadly neutralizing antibodies targeting conserved epitopes (CD4 binding site, MPER) that are poorly immunogenic in natural infection.
- Staphylococcal enterotoxins: Anti-toxin scFvs developed for both diagnostic lateral flow formats and passive immunotherapy in foodborne illness.
- Dengue virus: Cross-reactive antibodies against all four DENV serotypes selected using sequential panning against multiple antigen variants.
Next-Generation Advances in Phage Display Antibody Technology
NGS-Coupled Panning (HTPD)
High-throughput phage display (HTPD) integrates next-generation sequencing (NGS) of enriched pools after each round. Rather than picking individual colonies, the entire enriched library is sequenced, generating millions of reads. Enrichment ratios (read count round n÷round 1) identify convergent clones before ELISA screening even begins, dramatically compressing the screening funnel.
Machine Learning-Guided Library Design
- Deep learning models (e.g., AbLang, ESM-2) trained on Observed Antibody Space (OAS) databases predict functional VH/VL pairings for synthetic library design.
- Generative AI tools can design CDR3 sequences with predicted binding to a target from its structural coordinates alone.
- In silico affinity maturation using structure-based energy scoring reduces experimental optimization cycles.
Phage Display + Cryo-EM
Phage display hits are increasingly characterized by cryo-electron microscopy to map epitope locations at near-atomic resolution. This drives rational second-generation library design - a closed-loop where structural data feeds back into library construction.
Autonomous Robotic Panning Platforms
Fully automated liquid-handling systems (e.g., using Hamilton/Tecan robotics) now execute all panning rounds, washing, elution, re-infection, and phage ELISA in a single automated workflow, processing up to 8 targets in parallel within 3-4 weeks.
Conclusion
Phage display has fundamentally transformed antibody science. By coupling an antibody's phenotype (binding function) directly to its genotype (encoding DNA) within a single bacteriophage particle, it enables a selection process of extraordinary power - screening billions of antibody variants against any conceivable target in weeks, not years.
The technology's maturity is evidenced by over a dozen approved therapeutic antibodies, with adalimumab alone generating over $200 billion in cumulative sales. Beyond therapeutics, phage display antibody libraries are now the workhorse of research reagent development, enabling the sequence-defined, reproducible recombinant antibodies that modern scientific rigor demands.
As the field continues to integrate NGS-based deep screening, machine learning-guided library design, and robotic automation, phage display will remain indispensable, not just as an established antibody library discovery platform, but as an evolving technology at the intersection of molecular biology, data science, and precision medicine.
Faq's
What is phage display, and how does it work for antibody generation?
Phage display is an in vitro selection technique in which antibody genes (encoding scFv or Fab fragments) are fused to a bacteriophage coat protein gene. Each phage particle physically displays the encoded antibody on its surface while carrying the DNA inside. Libraries of billions of phage clones are incubated with a target antigen; non-binders are washed away, and binders are eluted, amplified, and subjected to further rounds of selection. After 3-5 rounds (biopanning), the enriched pool is screened individually, and positive clones are sequenced to identify the antibody.
What is the difference between a phage display antibody and a recombinant antibody?
All phage-display-derived antibodies are recombinant antibodies, meaning their variable domain sequences are expressed from defined recombinant DNA in a host system. The term "phage display antibody" describes the discovery method (in vitro biopanning), while "recombinant antibody" describes the production format (sequence-defined expression in a recombinant host). Recombinant antibodies can also come from other platforms, such as yeast display or ribosome display.
How large does a phage display antibody library need to be for drug discovery?
For research reagent discovery, libraries of 10⁷–10⁸ clones are often sufficient when using immune libraries against a known antigen. For therapeutic drug discovery targeting novel or difficult antigens, naïve or synthetic libraries with ≥10⁹ independent transformants are generally required. Some large synthetic libraries (e.g., HuCAL, Ylanthia) approach 10¹¹ functional clones, providing access to rare, high-affinity binders.
Can phage display generate fully human antibodies without animal immunization?
Yes. Naïve libraries constructed from B-cell repertoires of healthy, non-immunized human donors, or synthetic libraries built on human germline frameworks, yield fully human antibodies without any immunization step. This is one of the most compelling advantages of phage display over hybridoma technology, particularly for targets that are non-immunogenic or self-antigens in humans.
How many rounds of biopanning are typically needed in a phage display workflow?
Typically, 3-5 rounds are performed. Fewer rounds risk insufficient enrichment; more than 5 rounds risk over-selection toward clones with high phage infectivity rather than high antigen affinity. Counter-selection and increased wash stringency across rounds are more effective strategies than simply adding more rounds. NGS monitoring of enrichment kinetics helps determine the optimal stopping point.
What is the difference between scFv and Fab formats in phage display?
scFv (single-chain variable fragment) connects VH and VL with a peptide linker, making it compact and easy to clone, ideal for phage display and initial hit identification. Fab retains a more native antibody architecture (full heavy Fd + light chain), making it more stable and structurally representative for therapeutic development. Many workflows display scFv during selection, then reformat confirmed hits as full IgG for downstream characterization.
What are the main applications of phage display beyond antibody discovery?
Phage display has major applications in: (1) peptide library screening for receptor binding partners; (2) enzyme engineering via directed evolution; (3) vaccine development using phage particles as antigen carriers; (4) targeted drug delivery using phage-displayed peptides as tumor-homing ligands; (5) protein–protein interaction mapping; and (6) biosensor development for detection of small molecules, toxins, and pathogens.

 Cynthia
Cynthia